
Newsroom
The Cytidine-to-Uridine RNA editing is a significant post-transcriptional modification within organelles in higher plants, enriching the expression products of mitochondrial/ chloroplast genes. However, little is known about whether and to what extent RNA editing contributes to tissue-specific regulation from a global perspective.
A research team from the Wuhan Botanical Garden of the Chinese Academy of Sciences applied an integrative analysis of bioinformatics analysis of DNA-Seq and RNA-Seq data from roots and leaves of three tobacco varieties (TN90, Basma and K326) to identify RNA editing events and investigate the tissue specificity of RNA editing in plants.
According to the researchers, RNA editing was unevenly distributed between roots and leaves for all three varieties. The differentially edited sites located in plastid transcripts tended to occur in NDH (NADH dehydrogenase) complex subunits.
In general, the average RNA editing efficiency was obviously reduced in roots versus leaves, however, the reduction of RNA editing efficiency in mitochondria was mild compared with plastid.
The expression analysis of 22 differentially edited genes showed that 13 RNA-edited genes were down-regulated in roots with no genes up-regulated in roots (p value < 0.01), suggesting the expression and editing of transcripts were specifically regulated in different tissues.
Participated in recognizing edit sites, RNA editing factors, including pentatricopeptide repeat proteins and multiple organelle RNA editing factors proteins, showed an apparent down-regulated tendency in roots compared with leaves across three varieties in both organelles, which might limit the editing of sites in roots.
These findings indicate that the factor-mediated RNA editing is subject to tissue-dependent regulation, and the resultant organelle proteomes may be pertinent to the specific tissue functions.
Results have been published in Plant Biotechnology Reports with the title of "Tissuespecifcity of RNA editing in plant: analysis of transcripts from three tobacco (Nicotiana tabacum) varieties."
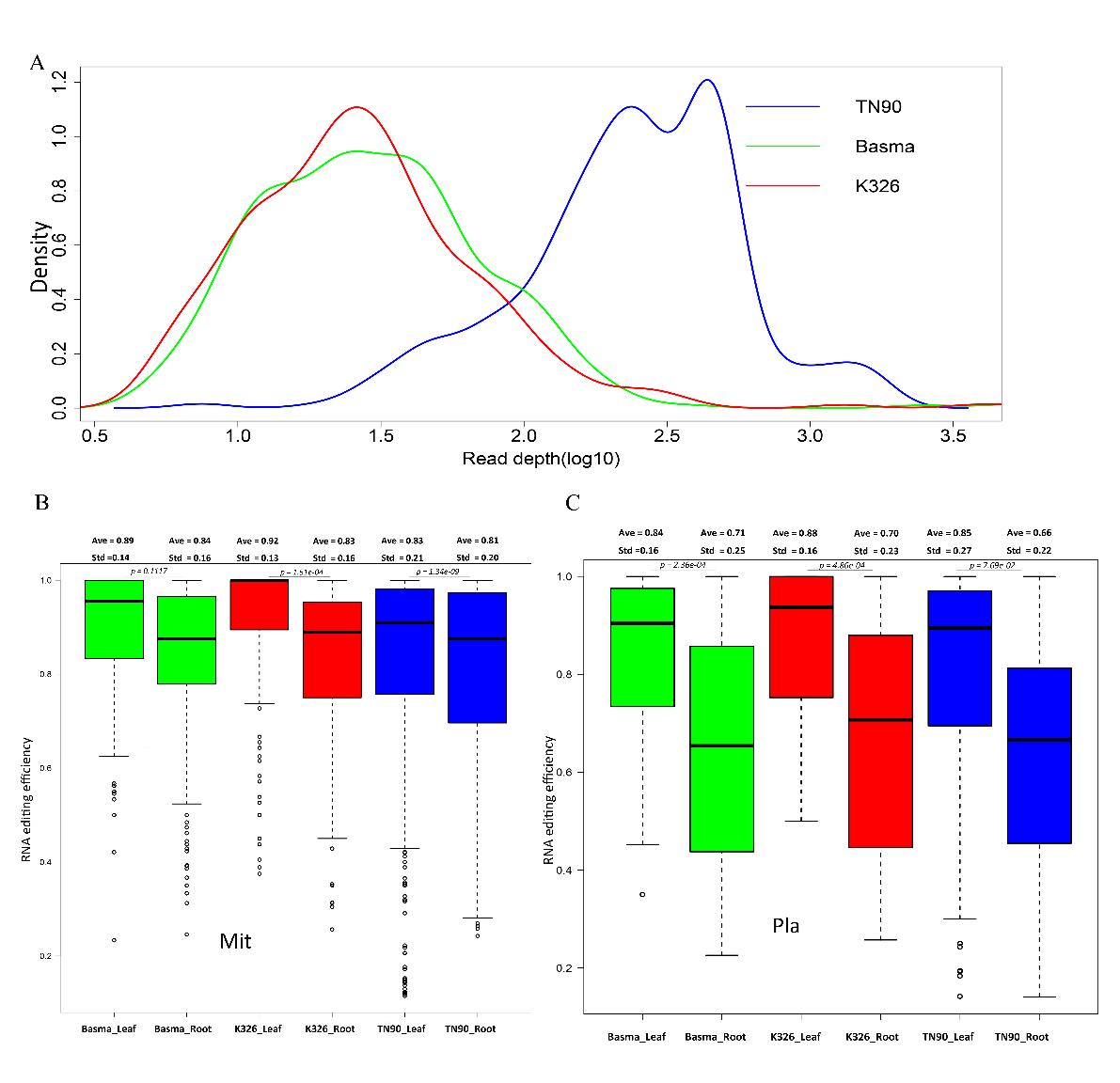
A. The overall density of RNA-seq read depth distribution of three tobacco varieties; B. RNA editing efficiency distribution of roots and leaves in mitochondria (Mit); C. RNA editing efficiency distribution of roots and leaves in plastid (Pla). (Image by WBG)
