
Based on big data integration and value-added curation, recently, the BIG Data Center at the Beijing Institute of Genomics of Chinese Academy of Sciences, constructed a series of featured databases, including EWAS Atlas (a knowledgebase of epigenome-wide association studies), iDog (an omics data resource for dog), EDK (Editome Disease Knowledgebase), PED (Plant Editosome Database), LncBook (a knowledgebase of human long non-coding RNAs), and NucMap (Nucleosome Positioning Map).
Seven articles were published online in the journal of Nucleic Acids Research (NAR), which will be released together in the NAR database issue in January 2019. Among them, six articles describe newly developed and updated databases and one provides an overview of database resources of the BIG Data Center.
In addition to these newly developed databases, the core database resources, namely, BioProject, BioSample, GSA, GWH, GVM, Science Wikis, and IC4R, have been significantly expanded and enriched. Also, several web services, viz., BIG Submission, BIG SSO, and BIG Search, have been significantly updated.
Especially, BIG Search has integrated data indexes from 20 partner databases, such as PlantTFDB, LncRNADisease, DEG, and lncRNASNP. To promote biodiversity and health big data sharing around the world, the BIG Data Center set up the Open Biodiversity and Health Big Data (BHBD) initiative.
As a major global center, the BIG Data Center has published its progresses in NAR annually since 2016, clearly demonstrating that the achievements of the BIG Data Center have been highly recognized by international counterparts. The BIG Data Center will continue to grow to offer a wide range of data services in support of worldwide research activities for big data deposition, integration and translation.
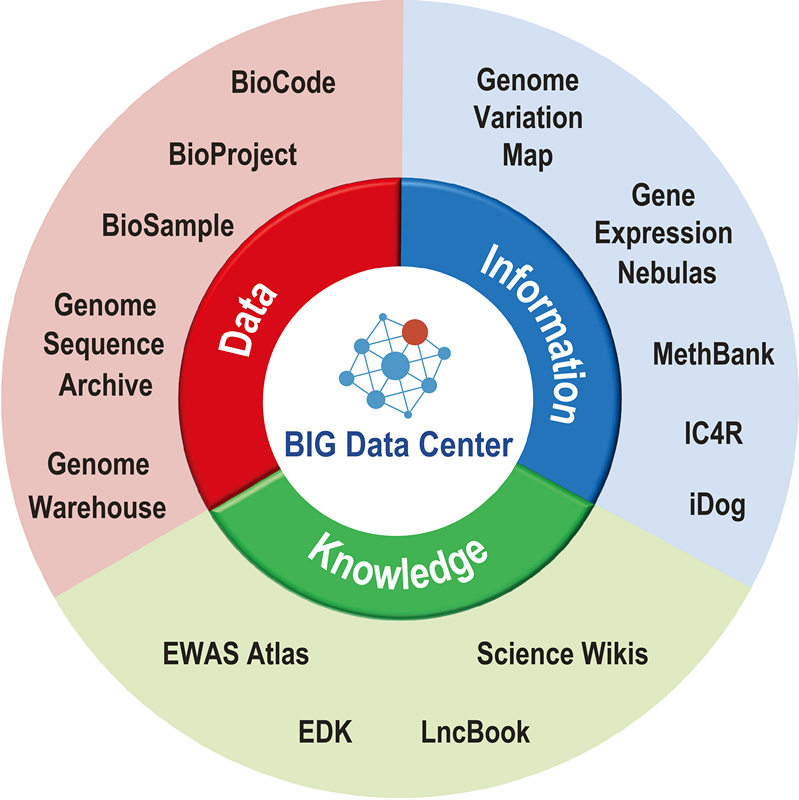
The BIG Data Center's core data resources (Image by BIG Data Center)
Article links:
Database Resources of the BIG Data Center in 2019
EWAS Atlas: a curated knowledgebase of epigenome-wide association studies
iDog: an integrated resource for domestic dogs and wild canids
Plant editosome database: a curated database of RNA editosome in plants
LncBook: a curated knowledgebase of human long non-coding RNAs
NucMap: a database of genome-wide nucleosome positioning map across species

86-10-68597521 (day)
86-10-68597289 (night)

52 Sanlihe Rd., Xicheng District,
Beijing, China (100864)

