
Newsroom
As a foundational benthic primary producer in aquatic ecosystems, Cladophora engages in complex interactions with associated bacteria that not only affect the equilibrium of these ecosystems but also drive the spread and evolution of antibiotic resistance genes (ARGs).
While previous studies have confirmed that the phycosphere serves as a critical niche for the occurrence and dissemination of ARGs in aquatic environments, a systematic understanding of how stage-dependent dissolved organic matter (DOM) release by Cladophora under natural conditions regulates the assembly of epiphytic phytoplankton and bacterial communities, the patterns of algae–bacteria interactions, and the subsequent dynamics of the associated resistome remains limited.
To address this knowledge gap, a research team led by Prof. WU Chenxi from the Institute of Hydrobiology (IHB) of the Chinese Academy of Sciences, has uncovered new insights into how life-stage-dependent processes within the macroalgal phycosphere control epiphytic community assembly and resistome dissemination in an oligotrophic lake. Their results highlight that Cladophora blooms function not only as modifiers of littoral biogeochemistry but also as important sources and amplifiers of antibiotic resistance risk.
The study was recently published in Water Research.
In this work, the researchers investigated Cladophora blooms in Qinghai Lake using a combination of field surveys and laboratory experiments. They examined how DOM derived from different algal morphotypes regulates the succession of epiphytic communities, with a specific focus on how shifts in community structure influence the spread of ARGs in the littoral zone.
Field observations showed that Cladophora blooms substantially modify littoral ecosystems by releasing DOM and nutrients, thereby shaping distinct epiphytic communities. The phycosphere harbors unique bacterial and phytoplankton assemblages characterized by lower diversity but greater functional specialization, dominated by taxa including Acinetobacter and Exiguobacterium. These microbes drive key nutrient cycles such as nitrogen metabolism and act as hotspots for resistomes. Contrary to the expectation that high cell density facilitates horizontal gene transfer (HGT), the results indicate that vertical gene transfer (VGT) is the primary mechanism underlying ARG proliferation in the phycosphere.
The team further explored shifts in bacterial community structure along a DOM gradient across phycospheres of different Cladophora morphotypes. They observed that under low-oxygen conditions, DOM composition transitions from labile, protein-like compounds to recalcitrant, humic-like substances. This transition promotes the enrichment of specific epiphytic communities marked by low diversity but high functional capacity. Moreover, bacterial diversity increased with decreasing DOM concentration along the gradient, underscoring the selective influence of algae on phycospheric bacteria.
This study establishes a mechanistic connection between algal life-stage-dependent DOM chemistry, selective microbial enrichment, and distinct evolutionary pathways of the resistome. It emphasizes the ecological risks posed by Cladophora blooms in facilitating the spread of resistance genes and disrupting biogeochemical cycling in littoral zones.
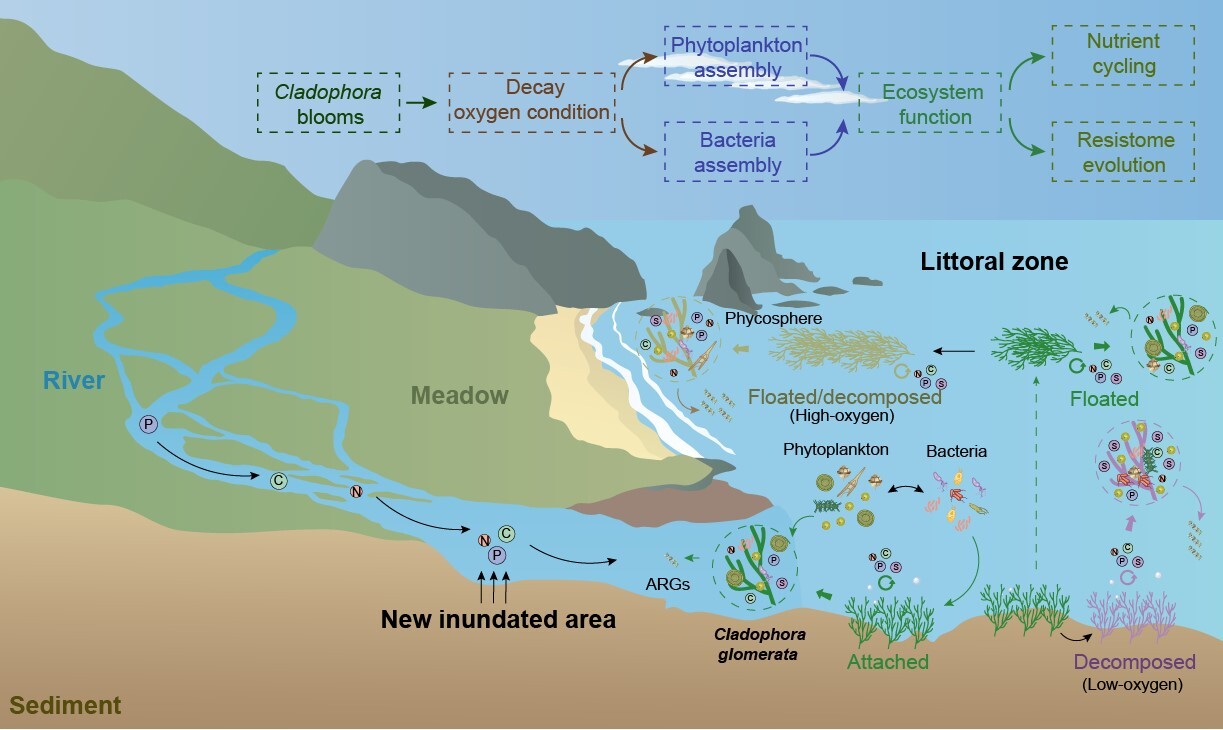
Effects of Cladophora blooms on nutrient cycling and antibiotic resistome evolution in the littoral zone of Qinghai Lake. (Image by IHB)
