
Nujiang River (NR), an essential component of the biodiversity hotspot of the Mountains of Southwest China, possesses a characteristic fish fauna and contains endemic species. Previous studies on fish diversity in the NR have primarily consisted of listings of the fish species observed during field collections, however, little is known about the genetic diversity of the NR fish fauna.
A research team led by Prof. HE Shunping from Institute of Hydrobiology, Chinese Academy of Sciences (IHB), combined long-term field work and molecular experiments to assess the genetic diversity of the NR fish species.
They collected and barcodes a total of 1,139 specimens belonging to 46 morphologically distinct fish species. According to their analyses, DNA barcoding is an efficient method for the identification of fish by the presence of barcode gaps. However, three invasive species are characterized by deep conspecific divergences, generating multiple lineages and Operational Taxonomic Units (OTUs), implying the possibility of cryptic species.
At the other end of the spectrum, ten species (from three genera) that are characterized by an overlap between their intra- and interspecific genetic distances form a single genetic cluster and share haplotypes. The neighbor-joining phenogram, Barcode Index Numbers (BINs) and Automatic Barcode Gap Discovery identified 43 putative species, while the General Mixed Yule-coalescence identified five more OTUs.
This study established a reliable DNA barcode reference library for the fish in the NR basin and sheds new light on the local fish diversity. Furthermore, this study could provide some important implications for protecting fish species, as well as other taxon in the NR basin. The research finding was published online in Scientific Reports journal entitled “The fish diversity in the upper reaches of the Salween River, Nujiang River, revealed by DNA barcoding”. This work was supported by the Key Fund and NSFC-Yunnan mutual funds of the National Natural Science Foundation of China.
 |
|
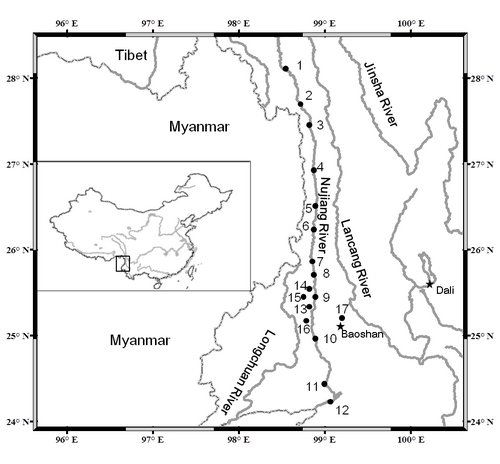
Map of locations sampled in this study. (Image by IHB) |
 |
| B: Haplotype networks of species with low interspecific divergences. (a) genusCreteuchiloglanis; (b) genusSchistura; and (c) genusSchizothorax.(Image by IHB) |

86-10-68597521 (day)
86-10-68597289 (night)

52 Sanlihe Rd., Xicheng District,
Beijing, China (100864)

