
Newsroom
Rice (Oryza sativa L.) is recently confirmed as a potential bioaccumulator plant of methylmercury (MeHg). Methylation of inorganic Hg influences the MeHg content in paddy soil, which directly affects the MeHg levels in rice seeds.
Studies on the mechanism of Hg methylation in rice paddies are significant to lower MeHg accumulation in rice and to reduce human health risks through contaminated rice consumption. MeHg cycling is mostly controlled by microbes but their importance in MeHg production and degradation in paddy soils remains unclear.
Dr. HU Haiyan and Dr. MENG Bo from the Institute of Geochemistry of the Chinese Academy of Sciences (IGCAS) revealed the roles of microbes in MeHg cycling in rice paddy soils. They employed a series of incubation experiments and stable isotope tracer techniques to investigate the relative importance of different microbial groups of MeHg production and degradation across a Hg contamination gradient (Fig.1).
The results showed that sulfate-reduction was the main driver of MeHg formation and concentration at non-contaminated sites. However, methanogenesis exhibited a complex and important role in controlling MeHg cycling at Hg mining sites.
The researchers further proposed that methanogenesis directly affected MeHg degradation via oxidative demethylation and indirectly affected MeHg production by out-competing other microbial guilds.
As a result, management of methanogenesis at Hg mining sites may shed new light on the potential for mitigation of MeHg production and reducing the risk of human exposure to MeHg.
Their findings were published in Environmental Science & Technology.
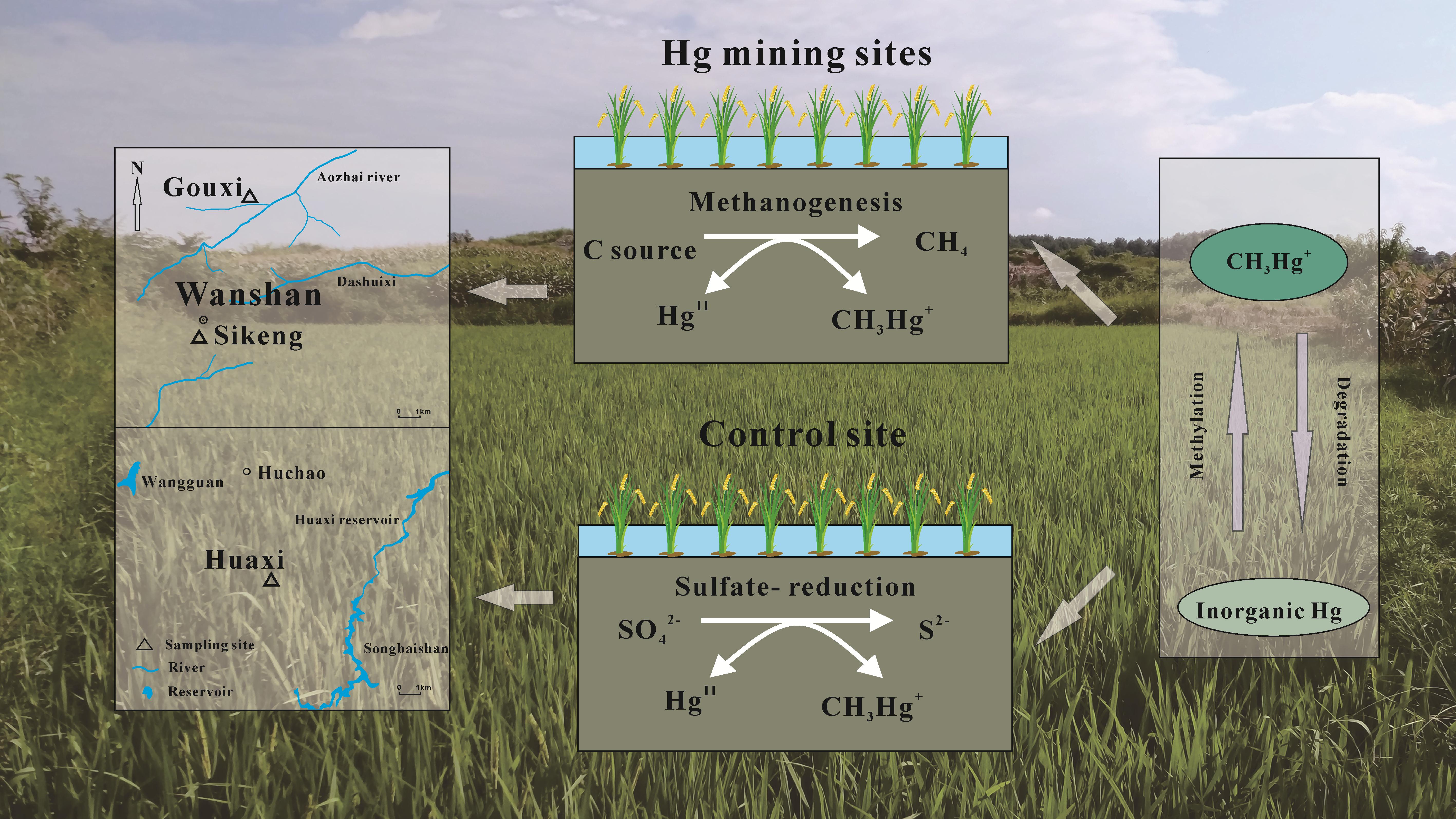
Fig. 1 Roles of microbes on methylmercury cycling in rice paddy soils (Image by IGCAS)
