The immune system operates with a critical baseline known as basal expression, or the "immune threshold." When this threshold is set too high, the plant risks autoimmune dysfunction; when set too low, it cannot mount a timely defense against pathogens.
In soybeans, which carry a vast repertoire of disease-resistance genes and a genome composed of more than 50% transposons, the mechanisms that govern this threshold have long remained elusive.
Long terminal repeat (LTR) retrotransposon is often dismissed as "selfish DNA" or genomic "fossils." A study published in
The Plant Cell and led by Dr. WANG Yuan’s group from the Institute of Genetics and Developmental Biology (IGDB) of the Chinese Academy of Sciences revealed that LTR retrotransposon are repurposed by the plant’s RNA-directed DNA methylation (RdDM) pathway to serve as a precise regulatory "dial" for immune gene expression.
Researchers identified an epigenetic control unit within the soybean genome. LTR transposon fragments were frequently positioned adjacent to key immune hubs, including TNL-type receptor genes and EDS1/SAG101 signaling components. Through RdDM pathway, the plant actively applied methylation marks to these transposons, maintaining adjacent defense genes in a low-level, "standby" state.
Then, researchers showed that when this epigenetic restraint was disrupted, nearby immune genes became constitutively activated even in the absence of pathogens, leading to widespread autoimmune activation and severely stunted plant growth.
The findings of this study demonstrate that the transposon module is essential for maintaining immune homeostasis, keeping large suites of defense genes poised at the ready while preventing harmful autoimmunity.
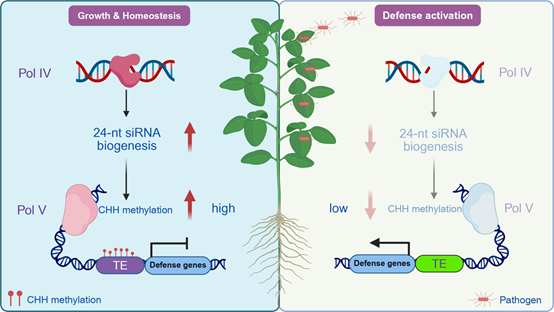
RdDM maintains the homeostasis of plant defense in soybean. (Image by IGDB)